|
||
|
About virus mentha
virus mentha archives evidence about viral interactions collected from different sources and presents these data in a complete and comprehensive way. Its data comes from manually curated protein-protein interaction databases that have adhered to the IMEx consortium. virus mentha is a resource that offers a series of tools to analyse selected proteins in the context of a network of interactions. Protein interaction databases archive protein-protein interaction (PPI) information from published articles. However, no database alone has sufficient literature coverage to offer a complete resource to investigate "the interactome". virus mentha's approach generates every week a consistent interactome (graph). Most importantly, the procedure assigns to each interaction a reliability score that takes into account all the supporting evidence. virus mentha offers direct access to viral families such as: Orthomyxoviridae, Orthoretrovirinae and Herpesviridae plus, it offers the unique possibility of searching by host organism. The website and the graphical application are designed to make the data stored in virus mentha accessible and analysable to all users. Go to User guide Our service displays content extracted from different sources. This content is responsibility of the entity that makes it available.
Reference
Please, in any article making use of the data extracted from VirusMentha, refer to: VirusMentha: a new resource for virus-host protein interactions Alberto Calderone, Luana Licata, Gianni Cesareni Nucleic Acids Res. 2014. doi:10.1093/nar/gku830 Data Integration Strategy and Data Source mentha: a resource for browsing integrated protein-interaction networks Alberto Calderone, Luisa Castagnoli, Gianni Cesareni Nature Methods 10, 690 (2013). doi:10.1038/nmeth.2561
Source databases 




mentha integrates these databases because of their characteristics. They all use the same curation policies and the same complex expansion model. Acknowledgment
User guideHow to search Search overview
Search example Search all elongation factors in arabidopsis thaliana
ALTERNATIVE
Result page
List page 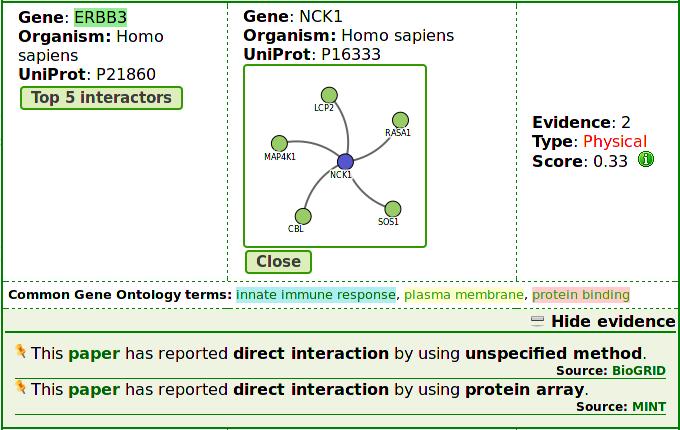
|
||
| University of Tor Vergata, Rome - Italy |



